Supported Interactions¶
Below is the full list of interactions currently supported by InterMap, including their geometric criteria and SMARTS definitions. Distance cutoffs are given in Angstroms (Å) and angles in degrees (°). All the reported values correspond to internal defaults and can be re-defined by users. Click on each interaction type to expand and view more details.
van der Waals¶
van der Waals contacts between any two atoms are detected if they are within the sum of their van der Waals radii. The van der Waals radii used are those defined by the RDKit library.
Close Contacts¶
Close contacts are defined as any two atoms that are within a specified threshold distance, typically set to 3.0 Å
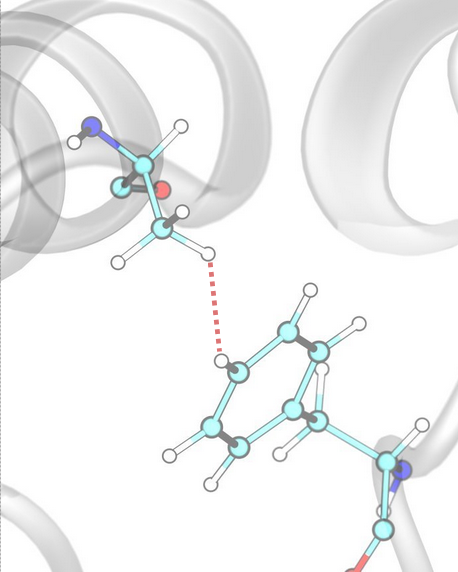
Hydrophobic¶
Hydrophobic interactions are defined based on the proximity of hydrophobic atoms, defined by the corresponding SMARTS pattern.
Hydrophobic Interactions
Cutoffs
- Distance
dist(i,j) <= 4.5
SMARTS
Hydrophobe: "[c,s,Br,I,S&H0&v2,$([C&R0;$([CH0](=*)=*),$([CH1](=*)-[!#1]),$([CH2](-[!#1])-[!#1])]),$([C;$([CH0](=*)(-[!#1])-[!#1]),$([CH1](-!#1])(-[!#1])-[!#1])]),$([C&D4!R](-[CH3])(-[CH3])-[CH3]);!$([#6]~[#7,#8,#9]);+0]"
Example
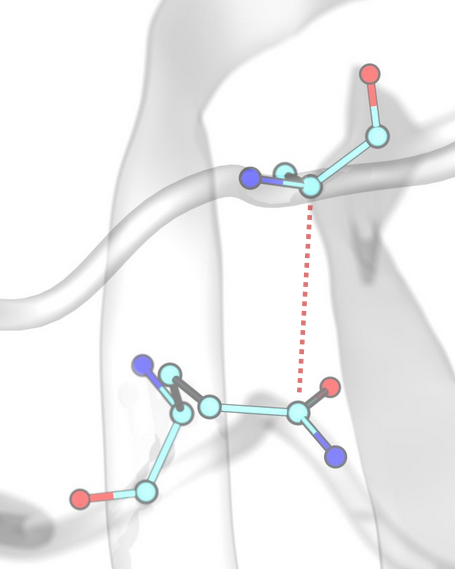
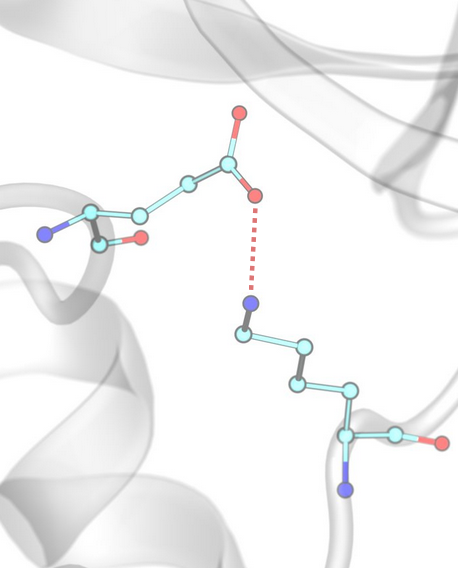
Anionic / Cationic (Salt Bridges)¶
Anionic and cationic interactions, commonly referred to as salt bridges, occur between oppositely charged groups, defined by the corresponding SMARTS patterns.
Anionic / Cationic (Salt Bridges) Interactions
Cutoffs
- Distance
dist(i,j) <= 4.5
SMARTS
Example
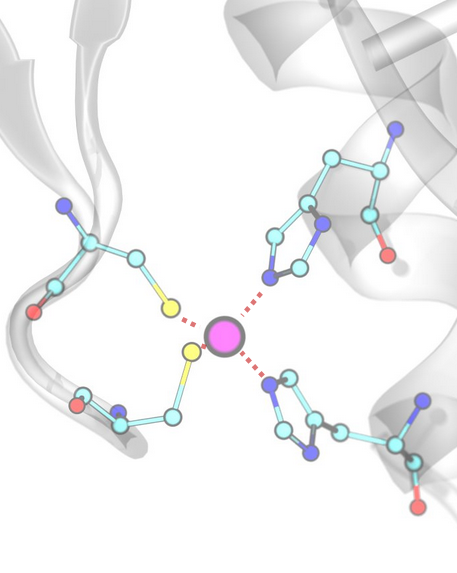
Metal Coordination¶
Interactions between metal ions and coordinating atoms, defined by the corresponding SMARTS patterns.
Metal Coordination
Cutoffs
- Distance
d(i,j) =< threshold(default:2.8 Å)
SMARTS
Metals: [Ca,Cd,Co,Cu,Fe,Mg,Mn,Ni,Zn]
Coordinating atoms: [O,#7&!$([nX3])&!$([NX3]-*=[!#6])&!$([NX3]-[a])&!$([NX4]),-{1-};!+{1-}]
Example
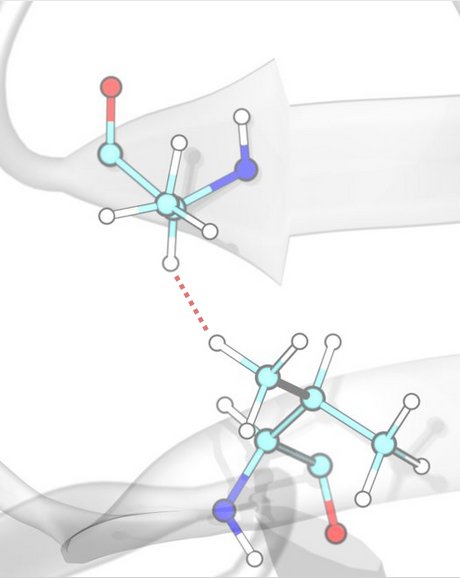
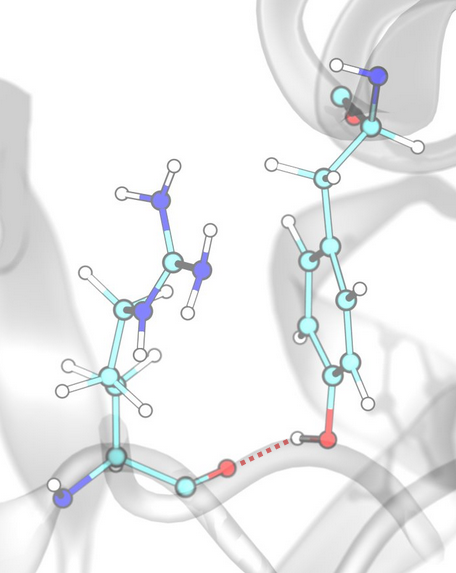
Hydrogen Bonds¶
Hydrogen bonds are detected based on distance and angle criteria between donor (D), hydrogen (H), and acceptor (A) atoms, defined by the corresponding SMARTS patterns of donors and acceptors.
Hydrogen Bond Interactions
Cutoffs
- Distance
dist(DA) <= 3.5 - Distance
dist(HA) <= 2.5 - Angle
130 <= ang(HD,HA) <= 180
SMARTS
Donor: [$([O,S,#7;+0]),$([Nv4+1]),$([n+]c[nH])]-[H]
Acceptor: [$([N&!$([NX3]-*=[O,N,P,S])&!$([NX3]-[a])&!$([Nv4+1])&!$(N=C(-[C,N])-N)]),$([n+0&!X3&!$([n&r5]:[n+&r5])]),$([O&!$([OX2](C)C=O)&!$(O(~a)~a)&!$(O=N-*)&!$([O-]-N=O)]),$([o+0]),$([F&$(F-[#6])&!$(F-[#6][F,Cl,Br,I])])]
Example
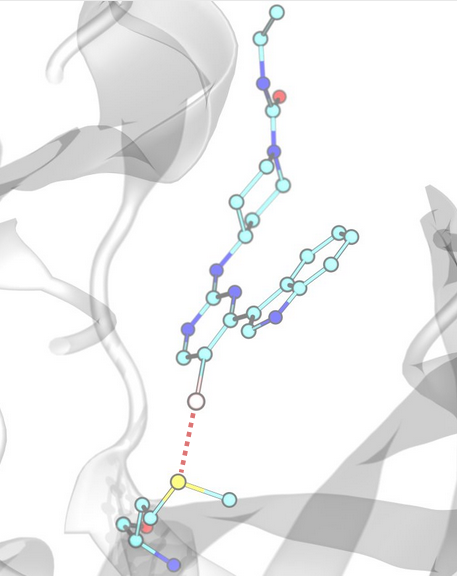
Halogen Bonds¶
Halogen bonds are detected based on distance and angle criteria between donors (D), halogen (X) and acceptors (A), defined by the corresponding SMARTS patterns of donors and acceptors.
Halogen Bond Interactions
Cutoffs
- Distance
dist(DA) <= 3.5 - Distance
dist(XA) <= 2.5 - Angle
130 <= ang(XD,XA) <= 180
SMARTS
Example
Cation–π / π-Cation¶
Cation–π interactions occur between a positively charged ion (cation) and the electron-rich π system of an aromatic ring. InterMaps detects these interactions based on distance criteria between the cation (C) and the centroid (R) of the aromatic ring, and the angle between the vector connecting the cation to the ring centroid (CR) and the normal to the ring plane (N).
Cation–π / π-Cation Interactions
Cutoffs
- Distance
dist(C,R) <= 4.5 - Angle
0 <= ang(CR, N) <= 30
SMARTS
Cation: [+{1-},$([NX3&!$([NX3]-O)]-[C]=[NX3+])]
Aromatic: [a;r6]1:[a;r6]:[a;r6]:[a;r6]:[a;r6]:[a;r6]:1
Example
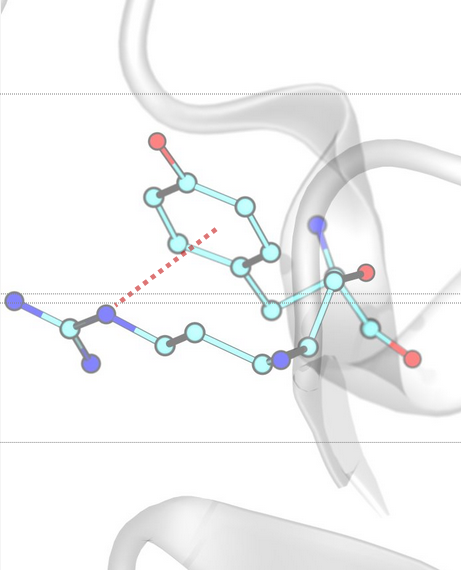
Anion–π / π-Anion¶
Anion–π interactions occur between a negatively charged ion (anion) and the electron-deficient π system of an aromatic ring. InterMaps detects these interactions based on distance criteria between the anion (A) and the centroid (R) of the aromatic ring, and the angle between the vector connecting the anion to the ring centroid (AR) and the normal to the ring plane (N).
Anion–π Interaction
Cutoffs
- Distance
dist(A,R) <= 4.5 - Angle
0 <= ang(AR, N) <= 30
SMARTS
Anion: [-{1-},$(O=[C,S,P]-[O-])]
Aromatic ring (6-members): [a;r6]1:[a;r6]:[a;r6]:[a;r6]:[a;r6]:[a;r6]:1
Aromatic ring (5-members): [a;r5]1:[a;r5]:[a;r5]:[a;r5]:[a;r5]:1
Example
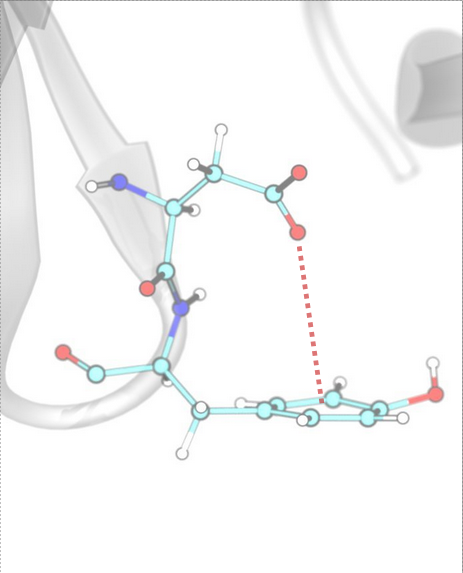
π–π Stacking [Face-to-Face]¶
Face-to-face π–π stacking interactions occur between two aromatic rings that are aligned parallel to each other. InterMaps detects these interactions based on distance criteria between the centroids of the rings (R1, R2) and the angle between their planes (N).
Face-to-Face π–π Stacking
Cutoffs
- Distance
dist(R1, R2) <= 5.5 - Angle
0 <= ang(N1, N2) <= 35 - Angle
0 <= ang(N1, R1R2) <= 30
SMARTS
Aromatic ring (6-members): [a;r6]1:[a;r6]:[a;r6]:[a;r6]:[a;r6]:[a;r6]:1
Aromatic ring (5-members): [a;r5]1:[a;r5]:[a;r5]:[a;r5]:[a;r5]:1
Example
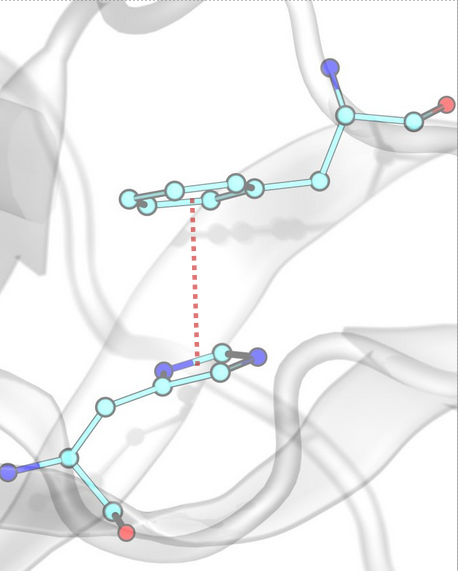
π–π Stacking [Edge-to-Face]¶
Edge-to-face π–π stacking interactions occur between two aromatic rings that are oriented perpendicularly to each other. InterMaps detects these interactions based on distance criteria between the centroids of the rings (R1, R2) and the angle between their planes (N).
Edge-to-Face π–π Stacking
Cutoffs
- Distance
dist(R1, R2) <= 6.5 - Angle
50 <= ang(N1, N2) <= 90
SMARTS
Aromatic ring (6-members): [a;r6]1:[a;r6]:[a;r6]:[a;r6]:[a;r6]:[a;r6]:1
Aromatic ring (5-members): [a;r5]1:[a;r5]:[a;r5]:[a;r5]:[a;r5]:1
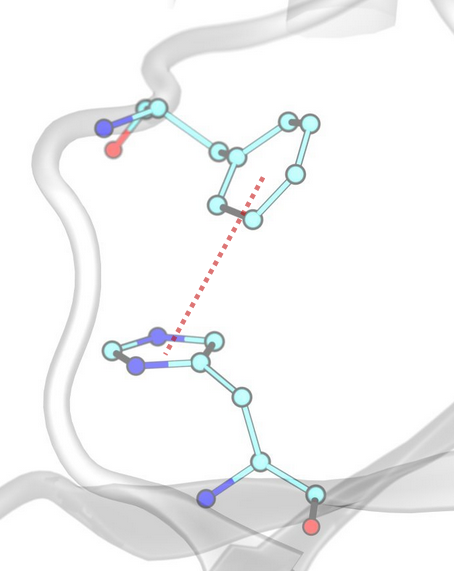
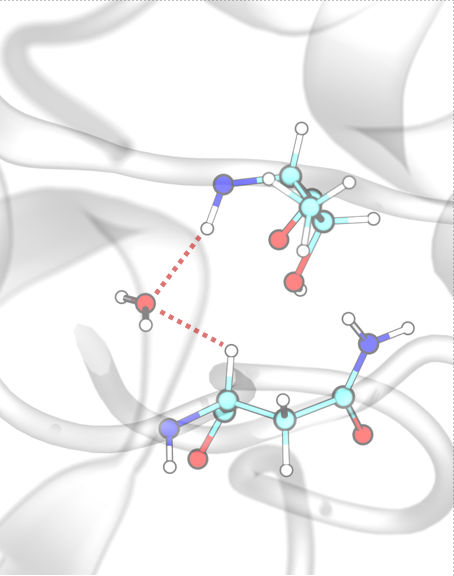
Water Bridges¶
InterMaps detects first-order water-mediated interactions (water bridges) when two residues form hydrogen bonds with the same water molecule. The criteria for hydrogen bond formation are the same as those defined for conventional hydrogen bonds.
First Order Water Bridges
SMARTS
Donor: [$([O,S,#7;+0]),$([Nv4+1]),$([n+]c[nH])]-[H]
Acceptor: [$([N&!$([NX3]-*=[O,N,P,S])&!$([NX3]-[a])&!$([Nv4+1])&!$(N=C(-[C,N])-N)]),$([n+0&!X3&!$([n&r5]:[n+&r5])]),$([O&!$([OX2](C)C=O)&!$(O(~a)~a)&!$(O=N-*)&!$([O-]-N=O)]),$([o+0]),$([F&$(F-[#6])&!$(F-[#6][F,Cl,Br,I])])]
Water: [O&H2]
Example